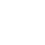RNASeqBrowser
Strand specific RNAseq data is now more common in RNAseq projects. The most widely used visualization tool is the UCSC genome browser that introduced the custom track concept that enabled researchers to simultaneously visualize gene expression at a particular locus from multiple experiments. This paper introduces a visualization tool (RNASeqBrowser) that incorporates and extends the functionality of the UCSC genome browser. For example, RNASeqBrowser simultaneously displays read coverage, SNPs, InDels and raw read tracks with other BED and wiggle tracks -- all being dynamically built from the BAM file. Paired reads are also connected in the browser to enable easier identification of novel exon/intron borders and chimaeric transcripts. Strand specific RNAseq data is also supported by RNASeqBrowser that displays reads above (positive strand transcript) or below (negative strand transcripts) a central line. Finally, RNASeqBrowser was designed for ease of use for users with few bioinformatic skills, and incorporates the features of many genome browsers into one platform.
Downloads
- standalone package and source code http://sourceforge.net/projects/rnaseqbrowser
- manual
- Launch the application with memory 1G , 3G , 5G, 12G(requires a good internet connection)
- Some browsers (chrome, safari) do not launch web start by default; in this case, double-click the downloaded jnlp file to execute it.
- If jnlp file is not opened by "java web start", set "java web start" as the default application for "jnlp" file. Alternatively, RNASeqBrowser can be launched from the command line by typing:
- >javaws http://data.australianprostatecentre.org/RNAseq/RNASeqBrowser/launch.jnlp
- >javaws http://data.australianprostatecentre.org/RNAseq/RNASeqBrowser/launch_3G.jnlp
- >javaws http://data.australianprostatecentre.org/RNAseq/RNASeqBrowser/launch_5G.jnlp
- >javaws http://data.australianprostatecentre.org/RNAseq/RNASeqBrowser/launch_12G.jnlp
- If an error "Java Application Blocked" appears, a new DeploymentRuleSet.jar file needs to be installed in the computer in the following directories, or simply remove the "Deployment" folder.
- On Windows platforms, install the file in the
<Windows-directory>\Sun\Java\Deploymentdirectory, for example,c:\Windows\Sun\Java\Deployment. - On Solaris or Linux platforms, install the file in the
/etc/.java/deploymentdirectory. - On Mac OS X platforms, install the file in the
/Library/Application Support/Oracle/Java/Deployment/directory.
- In Mac, if an error "can't be opened from an unidentified developer" appears, go to System Preferences -> Security & Privacy -> (Unlock Window) -> Allow apps downloaded from: (Select Anywhere). The procedure is shown in (figure).
- On Windows platforms, install the file in the
License
Permission to use this work is granted under the GNU Lesser General Public License (LGPL).
Enquiries
For enquiries about the software please contact Jiyuan An or Colleen Nelson.
Citing RNASeqBrowser
Jiyuan An, Lai J, Wood DLA, Sajjanhar A, Wang C, Tevz G, Lehman ML, Nelson CC, RNASeqBrowser: A genome browser for simultaneous visualization of raw strand specific RNAseq reads and UCSC genome browser custom tracks, BMC Genomics (in press). 2015