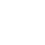J-Circos
Circos plots are graphical outputs that display three dimensional chromosomal interactions and fusion transcripts. However, the Circos plot tool is not an interactive visualization tool, but rather a figure generator. For example, it does not enable data to be added dynamically, nor does it provide information for specific data points interactively. Recently, an R-based Circos tool (RCircos) has been developed to integrate Circos to R, but similarly, Rcircos can only be used to generate plots. Thus, we have developed a Circos plot tool (J-Circos) that is an interactive visualization tool that can plot Circos figures, as well as being able to dynamically add data to the figure, and providing information for specific data points using mouse hover display and zoom in/out functions. J-Circos uses the Java computer language to enable it to be used on most operating systems (Windows, MacOS, Linux). Users can input data into J-Circos using flat data formats, as well as from the GUI. J-Circos will enable biologists to better study more complex chromosomal interactions and fusion transcripts that are otherwise difficult to visualize from next-generation sequencing data.
Downloads
- some browsers (chrome, safari) does not launch web start by default, you need to double click the downloaded jnlp file to run.
- In MacOS, if you get error "can't be opened from an unidentified developer" , System Preferences -> Security & Privacy -> (Unlock Window) -> Allow apps downloaded from: (Select Anywhere)
License
Permission to use this work is granted under the GNU Lesser General Public License (LGPL).
Enquiries
For enquiries about the software please contact Jiyuan An (j.an@qut.edu.au) or Colleen Nelson (colleen.nelson@qut.edu.au)
Tel: +61 7 3443 7280
Related tools
Circos, RCircos, ggbio, OmicCircos, circlize
Citing J-Circos
An J, Lai J, Sajjanhar A, Batra J, Wang C, Nelson CC (2014) J-Circos: An Interactive Circos plotter, Journal of Bioinformtics, DOI: 10.1093/bioinformatics/btu842